This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is transcriptomics?
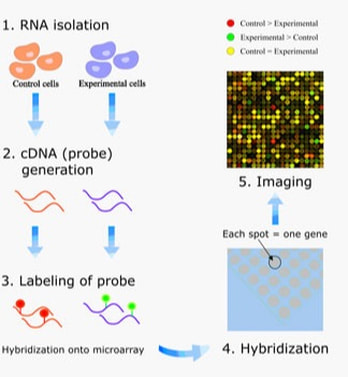
Transcriptomics is the study of the RNAs or transcripts found within a cell. Using an assay like a microarray (Figure 1) or other high-throughput sequencing methods, the specific RNA molecules found within the cell can be identified, thus providing valuable insight into which genes are being expressed [1]. In the case of myotonic dystrophy type 2 (DM2), transcriptomics can be used to determine which genes are up-regulated and down-regulated in the disease vs. non-disease state. This ultimately helps researchers determine what genes are being affected in DM2 individuals and can help identify any processes that may have become deregulated.
What is RNA sequencing?
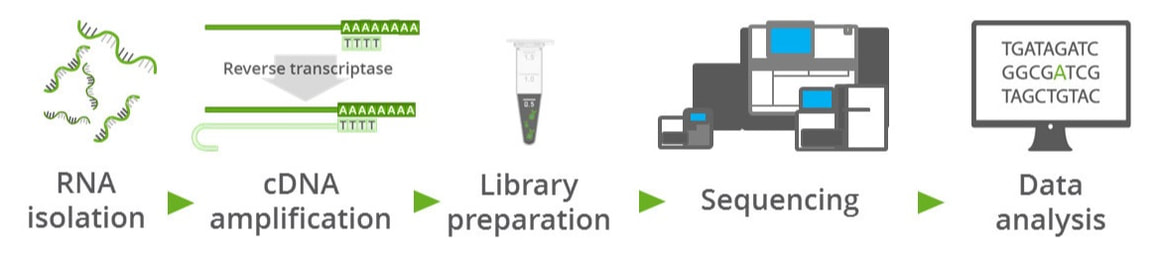
A fairly new technique that is beginning to replace assays like microarrays is RNA sequencing or RNA-seq (Figure 2), as it is commonly referred to. This technique allows for the quantification and discovery of mRNAs and other types of RNAs found within the cell. In this technique, RNAs are fragmented and converted to cDNA. Once in cDNA form, the bases can be sequenced using next-generation sequencing and the sequence of the original RNA transcript may be inferred [2]. This ultimately makes RNA-seq a very powerful technique in learning about what transcriptional changes may be occurring in a diseased cell vs. wild-type or control cell.
Discussion
In myotonic dystrophy type 2, CNBP pre-mRNA contains tetranucleotide repeats that cause the transcript to become toxic to the cell. As a result, an mRNA-gain-of-function mechanism can be seen, in which the pre-mRNA gains a detrimental function where it sequesters cellular proteins like MBNLs (see Gene Ontology page) [3].
Another consequence of this toxic pre-mRNA is the up-regulation of CUGBP1. This up regulation leads to detrimental splicing dysregulation throughout the cell. The total number of target mRNAs for CUGBP1 protein is unknown, however, thus constituting a gap in knowledge for DM2.
In addition to toxic CNBP pre-mRNA, CNBP protein levels have been shown to be diminished throughout the cell due to reduced translation. CNBP contains seven zinc-finger domains which are known to play a role in DNA and RNA binding, so low levels of CNBP in the disease state may lead to changes in the transcriptome [3].
Overall, transcriptomic research is a very viable future direction for DM2 research. By understanding what role toxic CNBP pre-mRNA plays within the cell, targeted mRNA gene therapies can be developed to eliminate the toxic pre-mRNAs.
Another consequence of this toxic pre-mRNA is the up-regulation of CUGBP1. This up regulation leads to detrimental splicing dysregulation throughout the cell. The total number of target mRNAs for CUGBP1 protein is unknown, however, thus constituting a gap in knowledge for DM2.
In addition to toxic CNBP pre-mRNA, CNBP protein levels have been shown to be diminished throughout the cell due to reduced translation. CNBP contains seven zinc-finger domains which are known to play a role in DNA and RNA binding, so low levels of CNBP in the disease state may lead to changes in the transcriptome [3].
Overall, transcriptomic research is a very viable future direction for DM2 research. By understanding what role toxic CNBP pre-mRNA plays within the cell, targeted mRNA gene therapies can be developed to eliminate the toxic pre-mRNAs.
References
- Transcriptomics. (n.d.). Retrieved March 16, 2018, from https://www.nature.com/subjects/transcriptomic
- An Introduction to RNA-seq. (2017, December 07). Retrieved from https://bitesizebio.com/13542/what-everyone-should-know-about-rna-seq/
- Meola, G., & Cardani, R. (2017). Myotonic dystrophy type 2 and modifier genes: An update on clinical and pathomolecular aspects. Neurological Sciences, 38(4), 535-546. doi:10.1007/s10072-016-2805-5
Header: https://i1.wp.com/www.artofthecell.com/wp-content/uploads/2014/10/John-Liebler-RNA-Polymerase.jpg?ssl=1