This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What are protein-protein interaction networks (PPIs)?
 Figure 1. Example of a PPI network.
Figure 1. Example of a PPI network.
The central dogma of biology is considered to be one of the most fundamental concepts in molecular biology and states that DNA codes for RNA, which is translated to proteins. Though most of the coding portions of the human genome eventually result in the formation of a single protein, these lone proteins do not act in act isolation. The interaction of various proteins within a cell constitutes the study of protein-protein interactions (PPIs). Proteins work together within cells to form accomplish biological processes and carry out molecular mechanisms. Some proteins, like serine/threonine kinases operate as part of signal-tranduction cascades, while other proteins, like CNBP, act as RNA and DNA binding proteins that influence the expression and translation of various genes [1,2].
Why are PPIs useful?
PPIs are useful because they help provide a global snapshot of how proteins are interacting within a cell. By understanding these various interactions, biological pathways can be teased out, and points of dysregulation can be pinpointed for diseases like myotonic dystrophy type 2 (DM2). DM2 is a complex disease that operates through various mechanisms, and understanding protein interactions is a vital step to identifying new areas of research and the underlying causes of DM2.
PPIs are important for understanding DM2
There are numerous online databases that provide information on protein-protein interactions including the String Database, IntAct, and BioGrid. These databases aggregate information on direct and indirect protein associations using predictive computational methodologies, translation of research in model organisms, and various other primary sources.
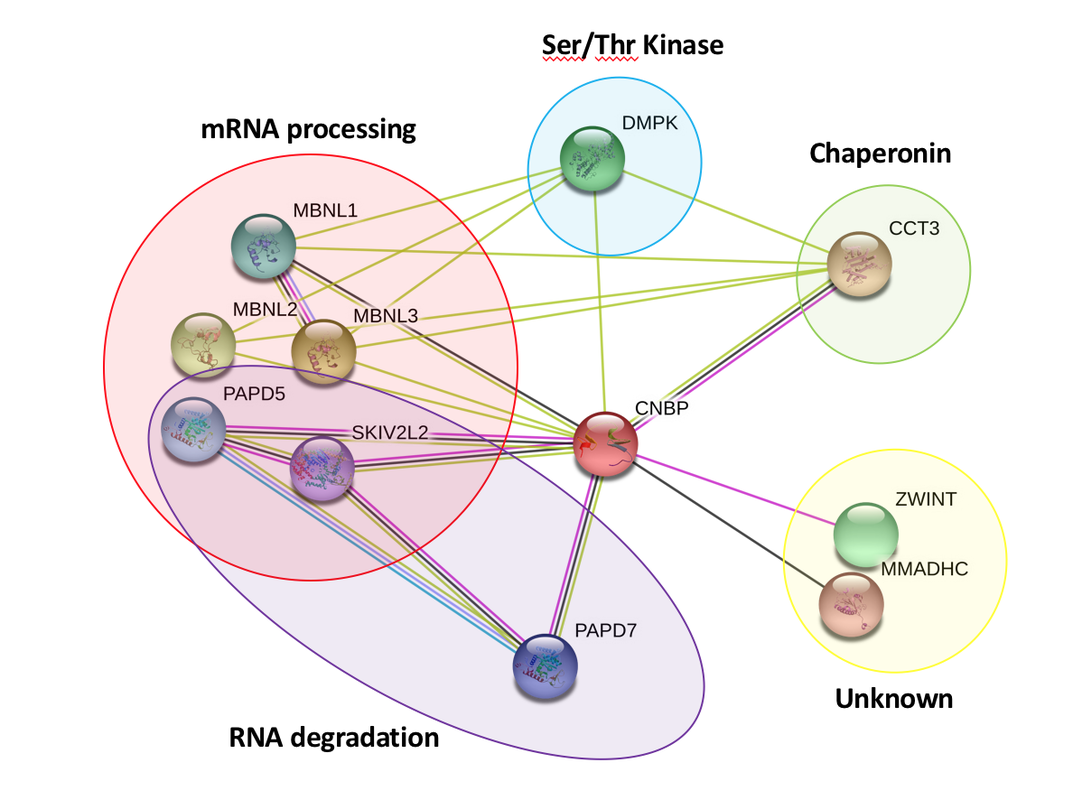
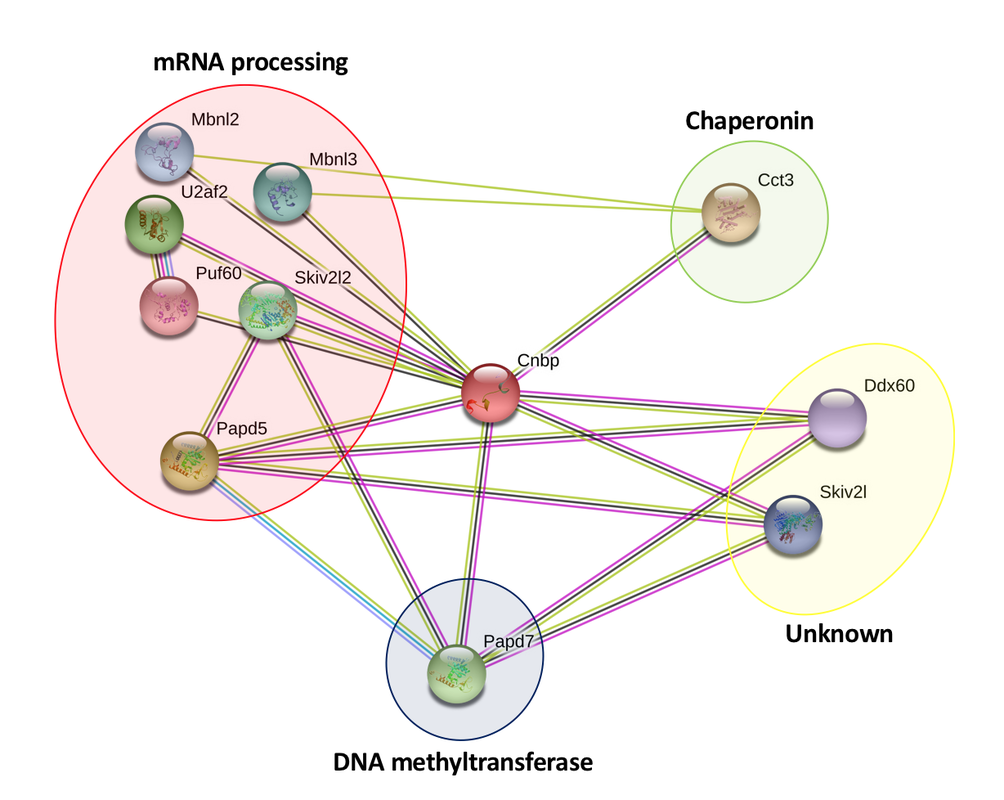
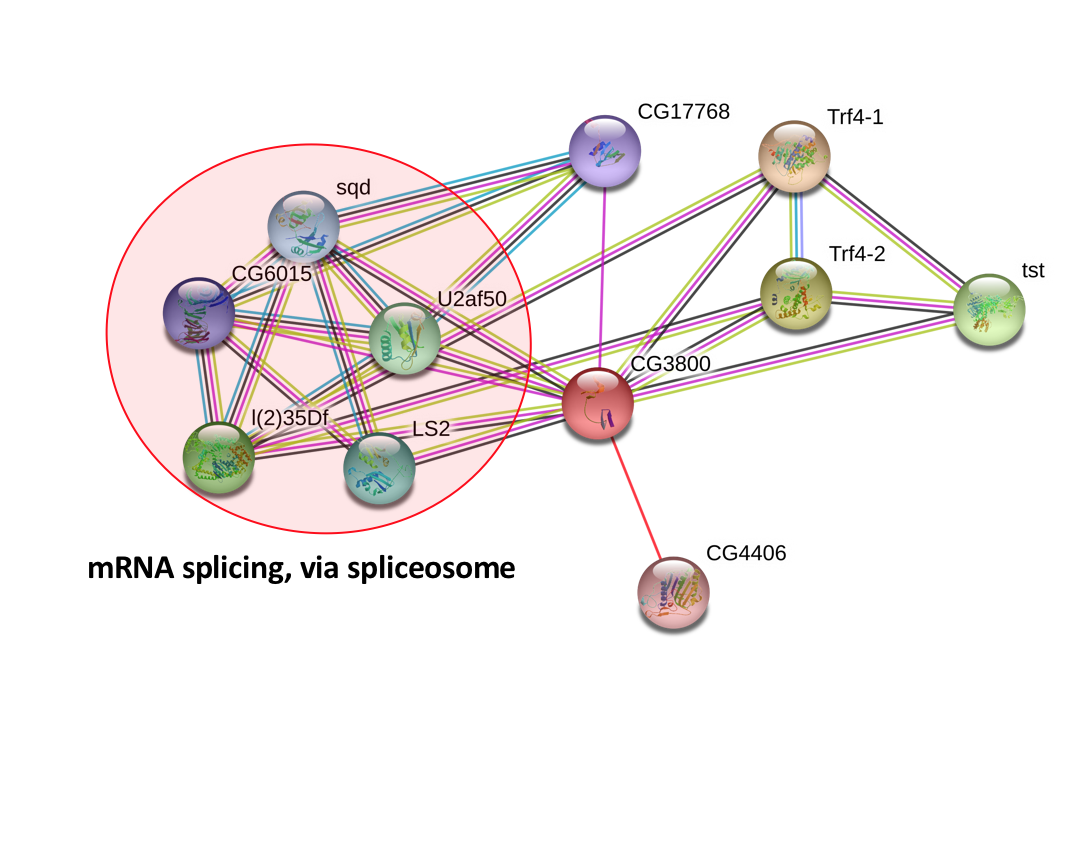
Using the String Database, CNBP interacting proteins were identified in Homo sapiens, Mus musculus, and Drosophila melanogaster (Figures 1, 2, & 3). Using PANTHER to confirm the gene ontology of various genes, it was found that many of the proteins found with the interaction network were involved in processes like mRNA processing, RNA degradation, and mRNA splicing. The colored, connecting lines also convey information about whether or not the interaction that appears in the diagram is a predicted or experimentally determined interaction.
Using the String Database, CNBP interacting proteins were identified in Homo sapiens, Mus musculus, and Drosophila melanogaster (Figures 1, 2, & 3). Using PANTHER to confirm the gene ontology of various genes, it was found that many of the proteins found with the interaction network were involved in processes like mRNA processing, RNA degradation, and mRNA splicing. The colored, connecting lines also convey information about whether or not the interaction that appears in the diagram is a predicted or experimentally determined interaction.
Discussion
The PPIs generated by the String Database convey information about the various proteins that interact with CNBP and its homologous proteins in other model organisms. From the diagram, it is clear that CNBP interacts with a variety of proteins, a large amount of which are involved in various RNA processes. Using these PPIs and continuing to find new proteins that fit within these networks will be critical to further understanding DM2. The role that decreased CNBP levels plays in human DM2 patients is still relatively unknown, and by building these networks, experiments that target interacting proteins can help researchers better understand how decreased CNBP is causing dysregulation of biological pathways that contribute to classic DM2 phenotypes like myotonia and muscle wasting.
References
- Protein-protein interaction networks. (2016, December 05). Retrieved from https://www.ebi.ac.uk/training/online/course/network-analysis-protein-interaction-data-introduction/protein-protein-interaction-networks
- CNBP CCHC-type zinc finger nucleic acid binding protein [Homo sapiens (human)] - Gene - NCBI. (n.d.). Retrieved from https://www.ncbi.nlm.nih.gov/gene/7555
Header: https://bmm.crick.ac.uk/~bmmadmin/Affinity/