This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is Homology?
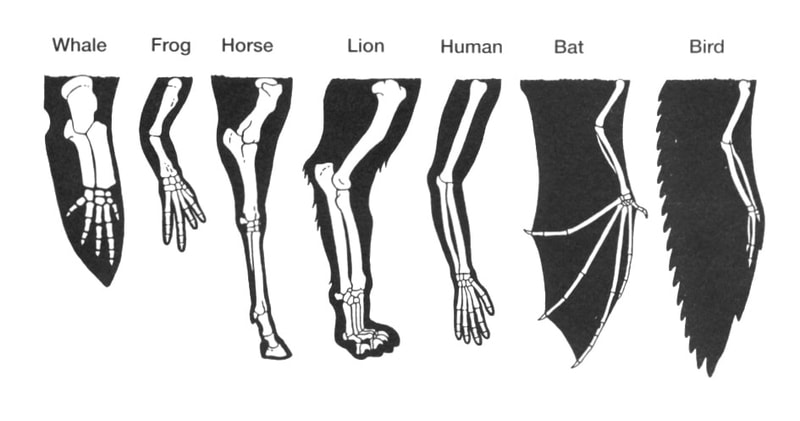
Homology refers to how similar the structural, developmental or genetic elements of one species are to another species. Similar to the relationships depicted in a phylogeny, homology is derived from organisms or species having a common evolutionary ancestor. As evolution takes places, bits and pieces of genetic information from past organisms can retain their structure and/or function. This is the principle of conservation. Conservation across species can take many forms, as in a particular gene having a similar nucleotide sequence amongst various species, or as in a limb with similar bone structure across species (Figure 1) [1].
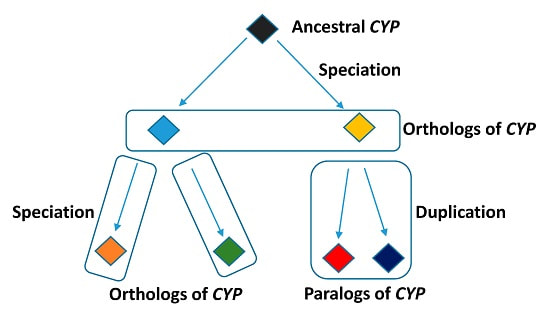
Homologs is a term that encompasses both orthogs and paralogs (Figure 2). Orthologs are genes found within different species that evolved from the same ancestral gene, and are genes that likely have similar functions across organisms. Paralogs are genes that are related by duplication events in an organism's genome. These duplicated genes may go on to evolve even more novel functions [2].
Homologs is a term that encompasses both orthogs and paralogs (Figure 2). Orthologs are genes found within different species that evolved from the same ancestral gene, and are genes that likely have similar functions across organisms. Paralogs are genes that are related by duplication events in an organism's genome. These duplicated genes may go on to evolve even more novel functions [2].
Homology of the CNBP Protein
Finding the homologs of a particular gene or protein in different species can be very beneficial for scientific research. By identifying genetic or proteomic homologs, appropriate model organisms for that gene or protein can be determined, and information about how that gene or protein functions in humans can be obtained. CNBP, for example, is a protein with a high degree of conservation amongst mammals. Shown below is a comparison of the percent identity or percent similarity of CNBP protein sequence data from NCBI's BLAST database across various species compared to the human form of CNBP that is relevant to myotonic dystrophy type 2.
Rattus norvegicus
|
Canis lupus familiaris
|
Gallus gallus
|
|
|
|
|
Danio rerio
|
Saccharomyces cerevisiae
|
Drosophila melanogaster
|
Discussion
CNBP is most highly conserved amongst mammals and muscular organisms, yet it is still fairly conserved in non-muscular single-cell organisms like yeast. The size of the protein ranges from 179 amino acids in humans to 153 amino acids in yeast. One surprising result of this analysis was that flies have a lower percent identity (36%) than yeast (41%), even though flies still contain small muscular structures. This suggests that CNBP may have other functions not related to muscle function.
References
- Britannica, T. E. (2016, September 08). Homology. Retrieved March 15, 2018, from https://www.britannica.com/science/homology-evolution
- Lewis, C. (n.d.). DEFINITION OF HOMOLOG, ORTHOLOG AND PARALOG. Retrieved March 15, 2018, from http://homepage.usask.ca/~ctl271/857/def_homolog.shtml
Header: https://www.thoughtco.com/analogous-homologous-structures-4079974